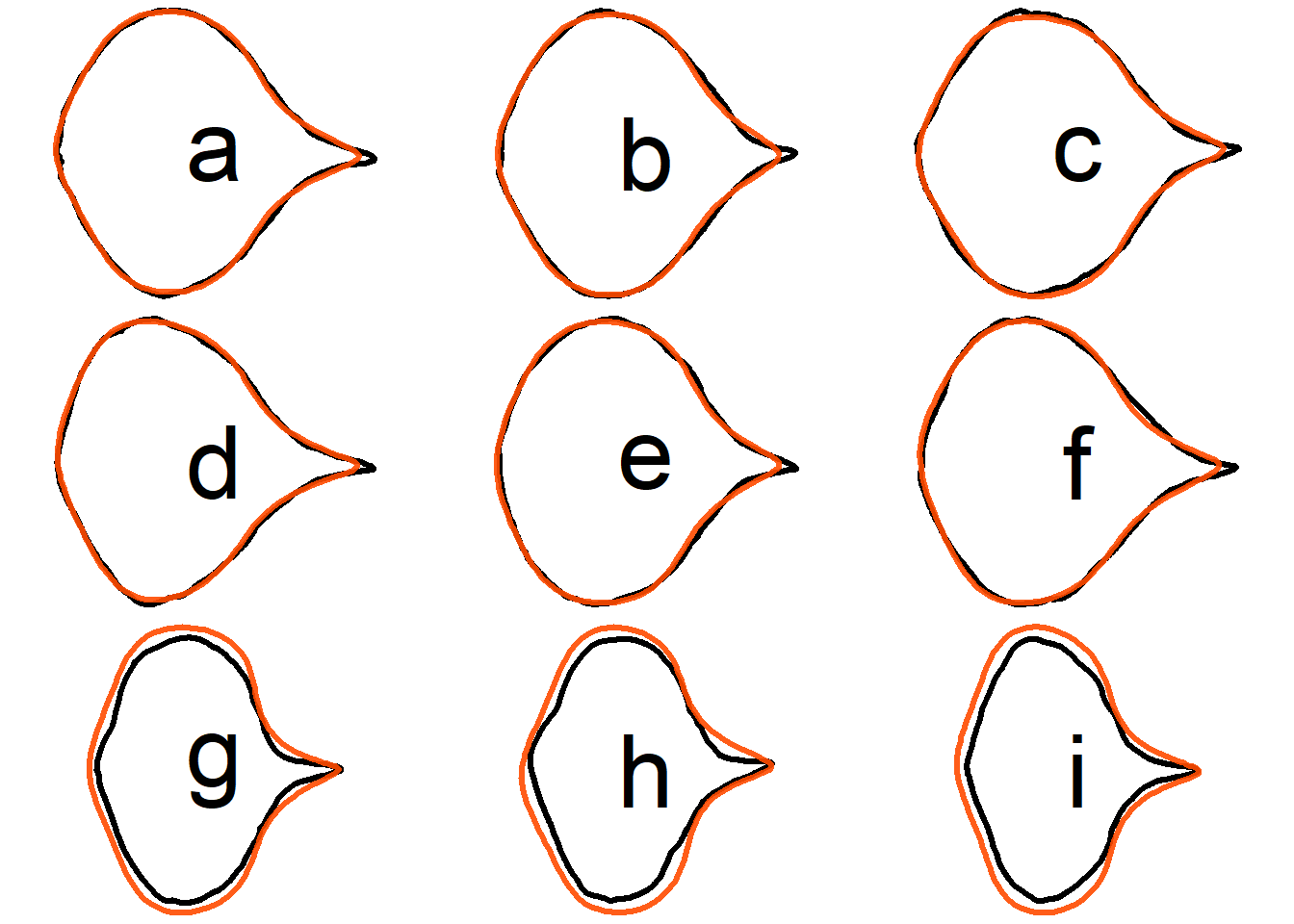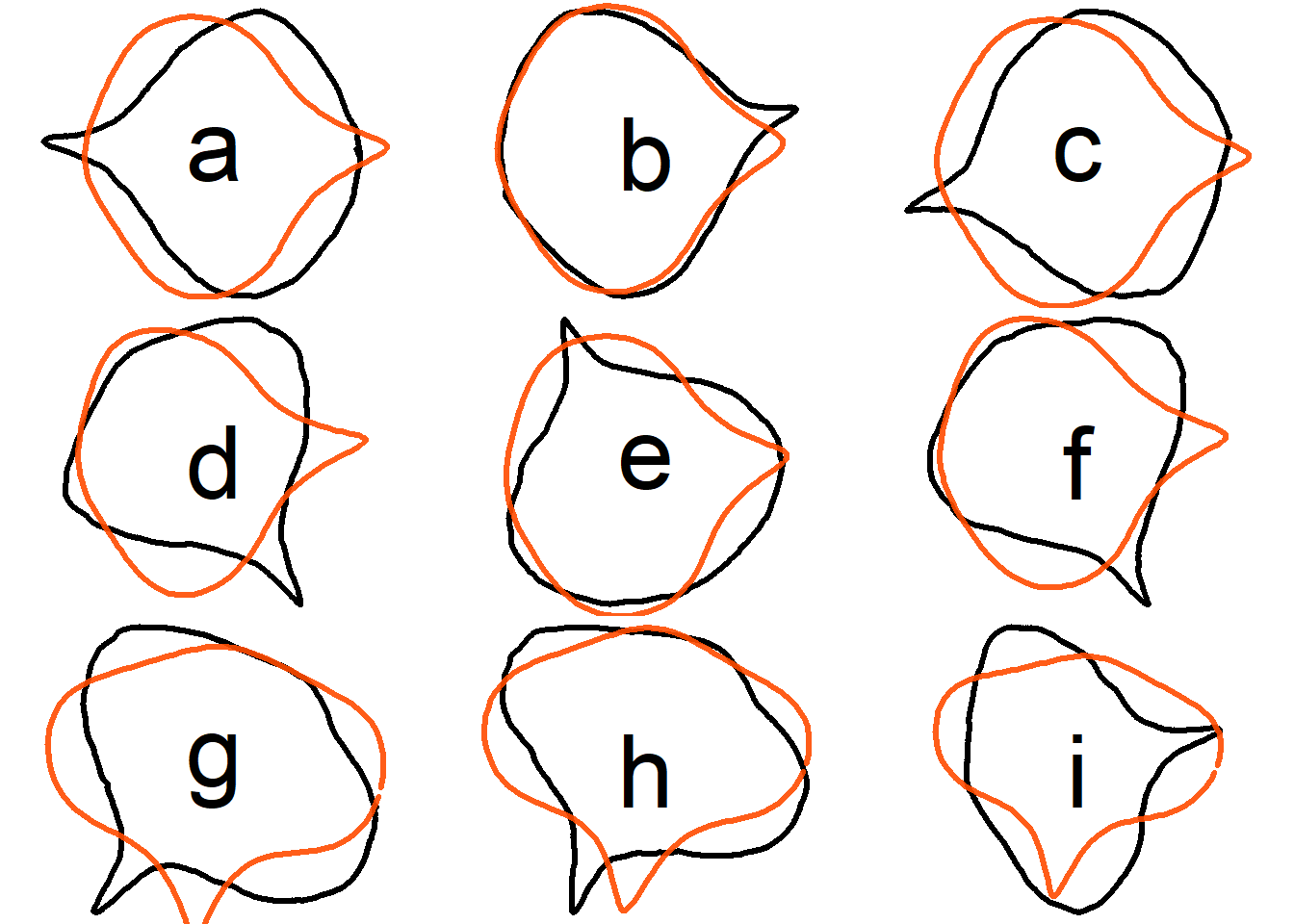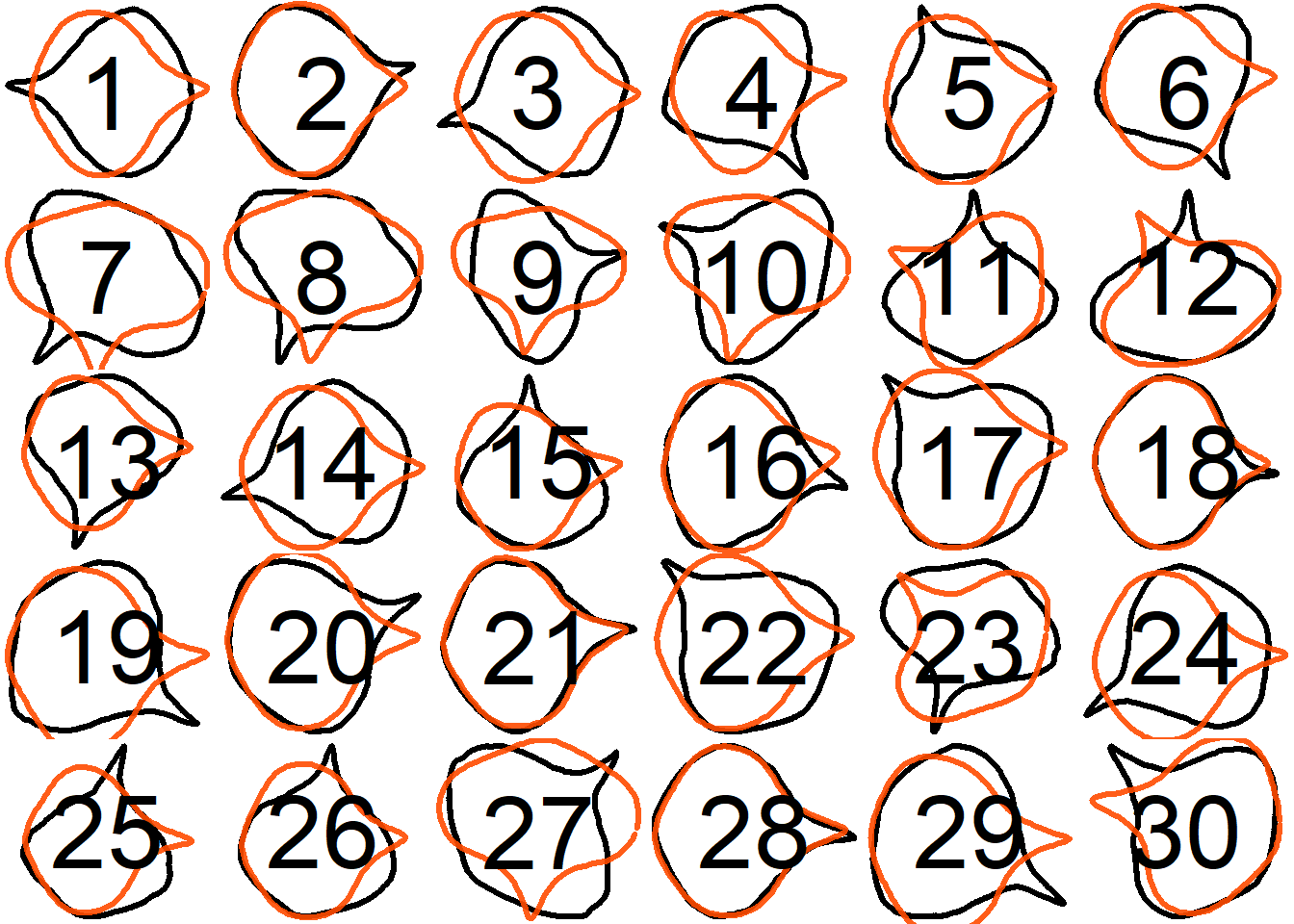---
title: "Direction check for *Triadica sebifera*"
---
This notebook compares original contours and contours reconstructed from oriented true EFD normalization and true EFD normalization for *Triadica sebifera*.
Output figures in this notebook are used in **Figure 3** of the manuscript.
## Setup
```{r}
#| label: setup
library(tools)
library(Momocs)
source("R/geometry.R")
source("R/normalization.R")
source("R/compare_contours.R")
palette("Okabe-Ito")
# set the data directory from environment variable (.Renviron)
data_dir <- Sys.getenv("PROJECT_DATA_DIR")
```
## Load contour data
Load the contour data from the CSV files in the `data/contour_triadica_sebifera/` directory.
```{r}
#| label: load_contours
file_paths_xy <- list.files(
file.path(data_dir, "data/contour_triadica_sebifera"),
full.names = TRUE,
pattern = "\\.csv$"
)
id_name <- file_path_sans_ext(basename(file_paths_xy))
xy_list <- lapply(file_paths_xy, function(fp) {
#cat("File:", fp, "\n")
df <- read.csv(fp)
df <- df[, c("x", "y")]
df <- as.matrix(df)
storage.mode(df) <- "double"
# if the contour is not closed, add the first point to the end to close it
if (!all(df[1, ] == df[nrow(df), ])) {
df <- rbind(df, df[1, ])
}
return(df)
})
names(xy_list) <- id_name
```
## Load oriented true normalized EFD coefficients
True normalized EFD coefficients data are obtained by [LeafContourEFD](https://github.com/maple60/leaf-contour-efd).
```{r}
#| label: load_oriented_true_normalized_coefficients
file_paths <- list.files(
file.path(data_dir, "data/coefficients_efd_normalized_triadica_sebifera"),
full.names = TRUE,
pattern = "\\.csv$"
)
ef_list <- lapply(file_paths, function(fp) {
#cat("File:", fp, "\n")
df <- read.csv(fp)
return(df)
})
```
## Oriented true EFD normalization
Compare the original contour and the reconstructed contour from **oriented true EFD normalization** for the first 9 samples.
```{r}
#| label: compare_oriented_true_EFD_normalization_examples
extract_id <- 1:9
labels <- letters[seq_along(extract_id)]
layout(matrix(seq_along(extract_id), nrow = 3, byrow = TRUE))
for (i in seq_along(extract_id)) {
compare_contour_oriented_true_EFD_normalization(
original = xy_list[[extract_id[i]]],
ef_normalized = ef_list[[i]],
nb.h = 5,
mar = c(0, 0, 0, 0),
lwd = 3,
cex.text = 5,
label = labels[i]
)
}
```
```{r}
#| label: reconstruct_contour_oriented_true_EFD_normalization
#| eval: false
#| code-fold: true
#| code-summary: "Save the plot as SVG file"
save_dir <- "results"
dir.create(save_dir, showWarnings = FALSE)
save_file_name <- "fig3a_compare_contours_oriented_true_efd_normalized_triadica_sebifera.svg"
svg(file.path(save_dir, save_file_name), bg = "transparent")
extract_id <- 1:9
labels <- letters[seq_along(extract_id)]
layout(matrix(seq_along(extract_id), nrow = 3, byrow = TRUE))
for (i in seq_along(extract_id)) {
compare_contour_oriented_true_EFD_normalization(
original = xy_list[[extract_id[i]]],
ef_normalized = ef_list[[i]],
nb.h = 5,
mar = c(0, 0, 0, 0),
lwd = 3,
cex.text = 5,
label = labels[i]
)
}
dev.off()
```
## True EFD normalization
To assess whether contours reconstructed after **true EFD normalization** are oriented consistently with the original contours, we randomly rotate the original contours before applying **true EFD normalization**.
```{r}
#| label: rotate_contours
set.seed(123) # set.seed for reproducibility
# rotation angles in degrees
angles <- sample(0:360, length(xy_list), replace = TRUE)
# rotate contours
xy_list_rotated <- mapply(
function(mat, ang) rotate_xy_centered(mat, ang),
xy_list,
angles,
SIMPLIFY = FALSE
)
```
Compare the original contour and the reconstructed contour from **true EFD normalization** for the first 9 samples.
```{r}
#| label: compare_true_EFD_normalization_examples
xy_list_rotated_example <- xy_list_rotated[1:9]
labels <- letters[seq_along(xy_list_rotated_example)]
layout(matrix(seq_along(xy_list_rotated_example), nrow = 3, byrow = TRUE))
for (i in seq_along(xy_list_rotated_example)) {
compare_contour_true_EFD_normalization(
original = xy_list_rotated_example[[i]],
nb.h = 5,
mar = c(0, 0, 0, 0),
lwd = 3,
cex.text = 5,
label = labels[i]
)
}
```
```{r}
#| label: reconstruct_contour_true_EFD_normalization
#| eval: false
#| code-fold: true
#| code-summary: "Save the plot as SVG file"
save_dir <- "results"
dir.create(save_dir, showWarnings = FALSE)
save_file_name <- "fig3b_compare_contours_true_efd_normalized_triadica_sebifera.svg"
svg(file.path(save_dir, save_file_name), bg = "transparent")
xy_list_rotated_example <- xy_list_rotated[1:9]
labels <- letters[seq_along(xy_list_rotated_example)]
layout(matrix(seq_along(xy_list_rotated_example), nrow = 3, byrow = TRUE))
for (i in seq_along(xy_list_rotated_example)) {
compare_contour_true_EFD_normalization(
original = xy_list_rotated_example[[i]],
nb.h = 5,
mar = c(0, 0, 0, 0),
lwd = 3,
cex.text = 5,
label = labels[i]
)
}
dev.off()
```
## Supplementary figures
Visualize all the original contours and the contours reconstructed from oriented true EFD normalization for all samples, and save the plots as SVG files.
```{r}
#| label: reconstruct_contour_oriented_true_EFD_normalization_all
layout(matrix(seq_along(xy_list_rotated), nrow = 5, byrow = TRUE))
for (i in seq_along(xy_list_rotated)) {
compare_contour_true_EFD_normalization(
original = xy_list_rotated[[i]],
nb.h = 5,
mar = c(0, 0, 0, 0),
lwd = 3,
cex.text = 5,
label = i
)
}
```
```{r}
#| label: save_comparison_plots_all
#| eval: false
#| code-fold: true
#| code-summary: "Save the plot as SVG file"
save_dir <- "results"
dir.create(save_dir, showWarnings = FALSE)
save_file_name <- "supplementary_compare_contours_true_efd_normalized_all_triadica_sebifera.svg"
svg(file.path(save_dir, save_file_name), bg = "transparent")
layout(matrix(seq_along(xy_list_rotated), nrow = 5, byrow = TRUE))
for (i in seq_along(xy_list_rotated)) {
compare_contour_true_EFD_normalization(
original = xy_list_rotated[[i]],
nb.h = 5,
mar = c(0, 0, 0, 0),
lwd = 3,
cex.text = 5,
label = i
)
}
dev.off()
```