---
title: "PCA of *Quercus serrata* and *Quercus crispula* leaf contours"
---
The goals of this notebook are:\
1. To perform PCA using the coefficients obtained by conventional normalization, oriented true EFD normalization and normalized EFD.\
2. To compare PCA results across these three methods.
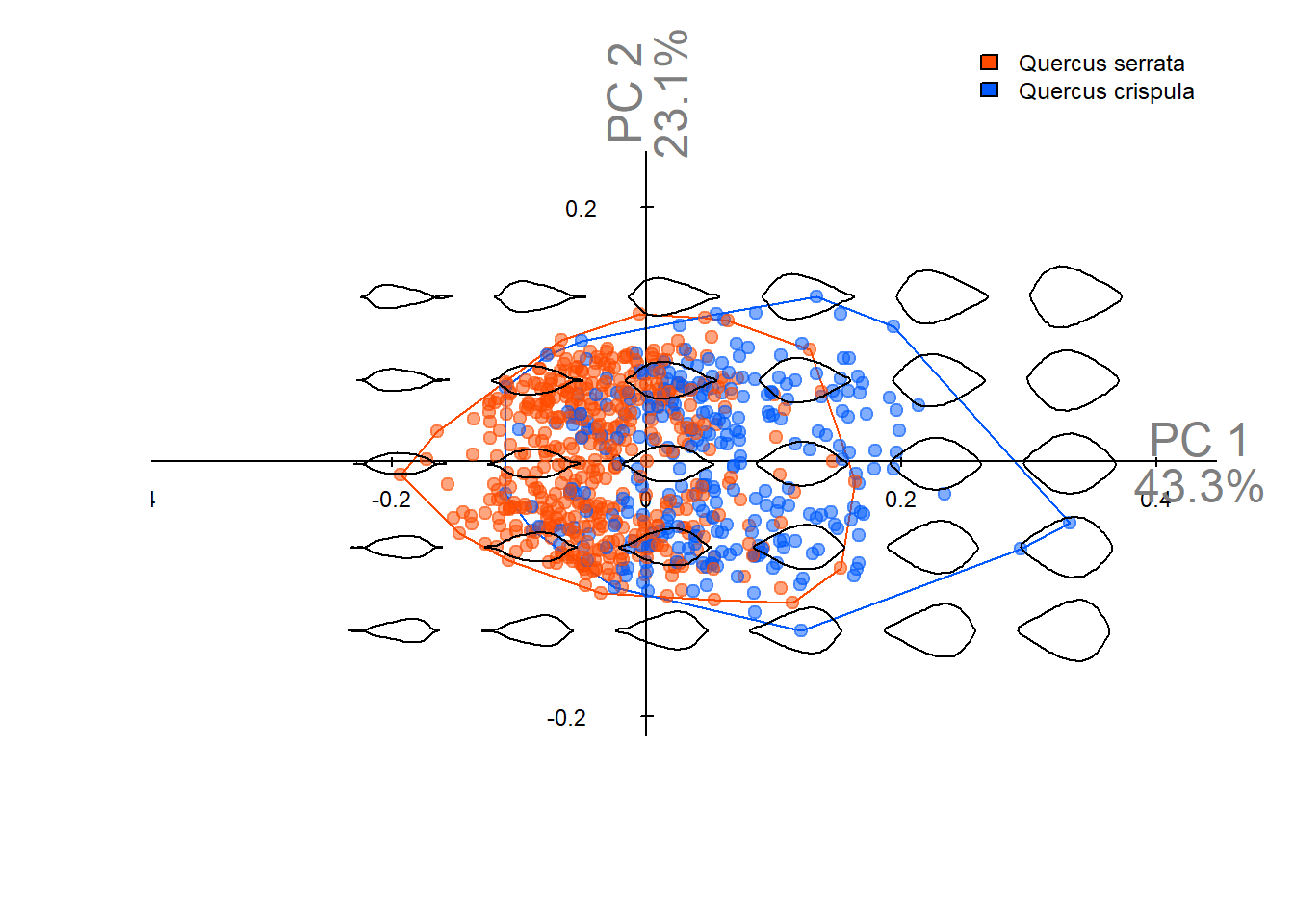
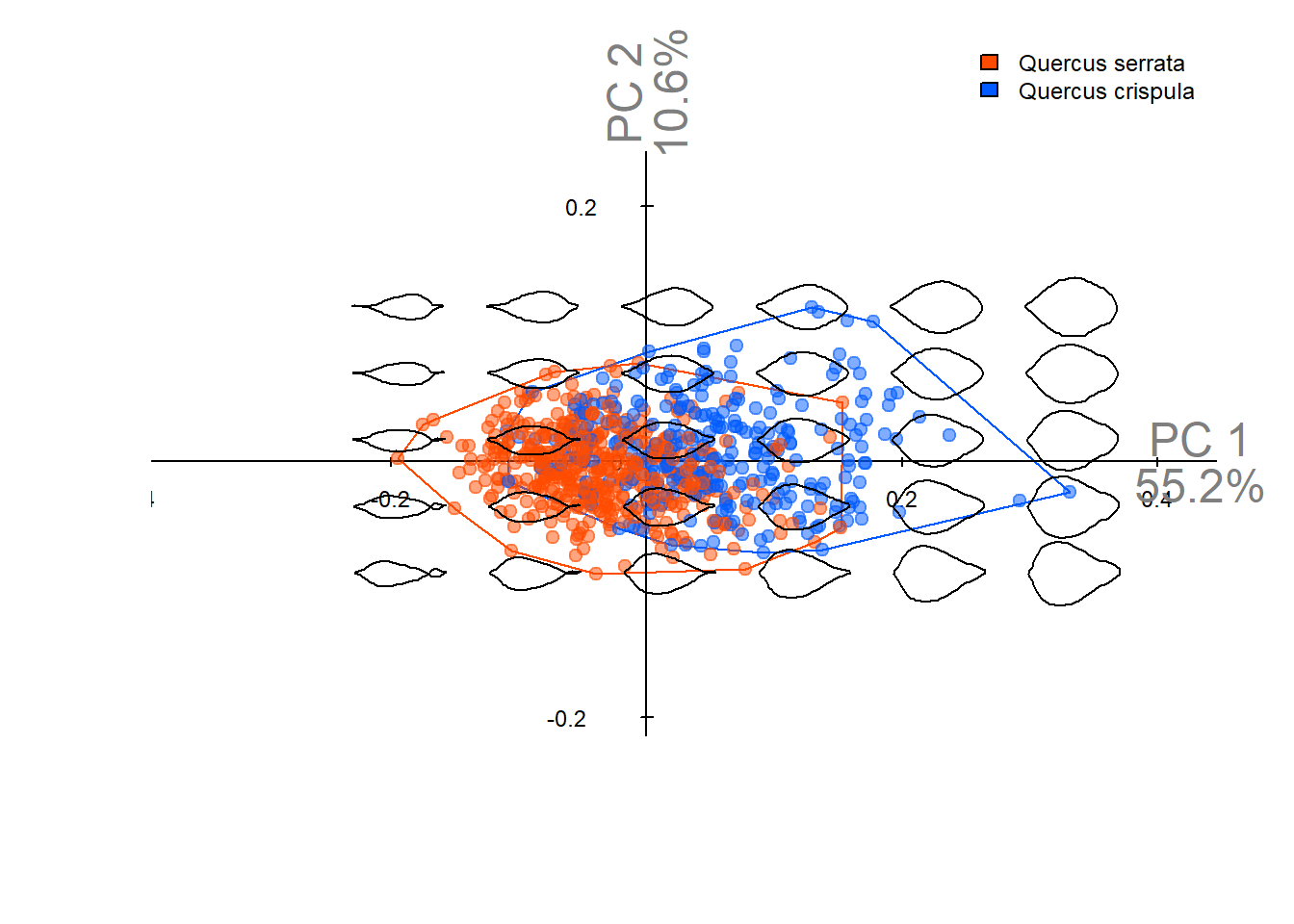
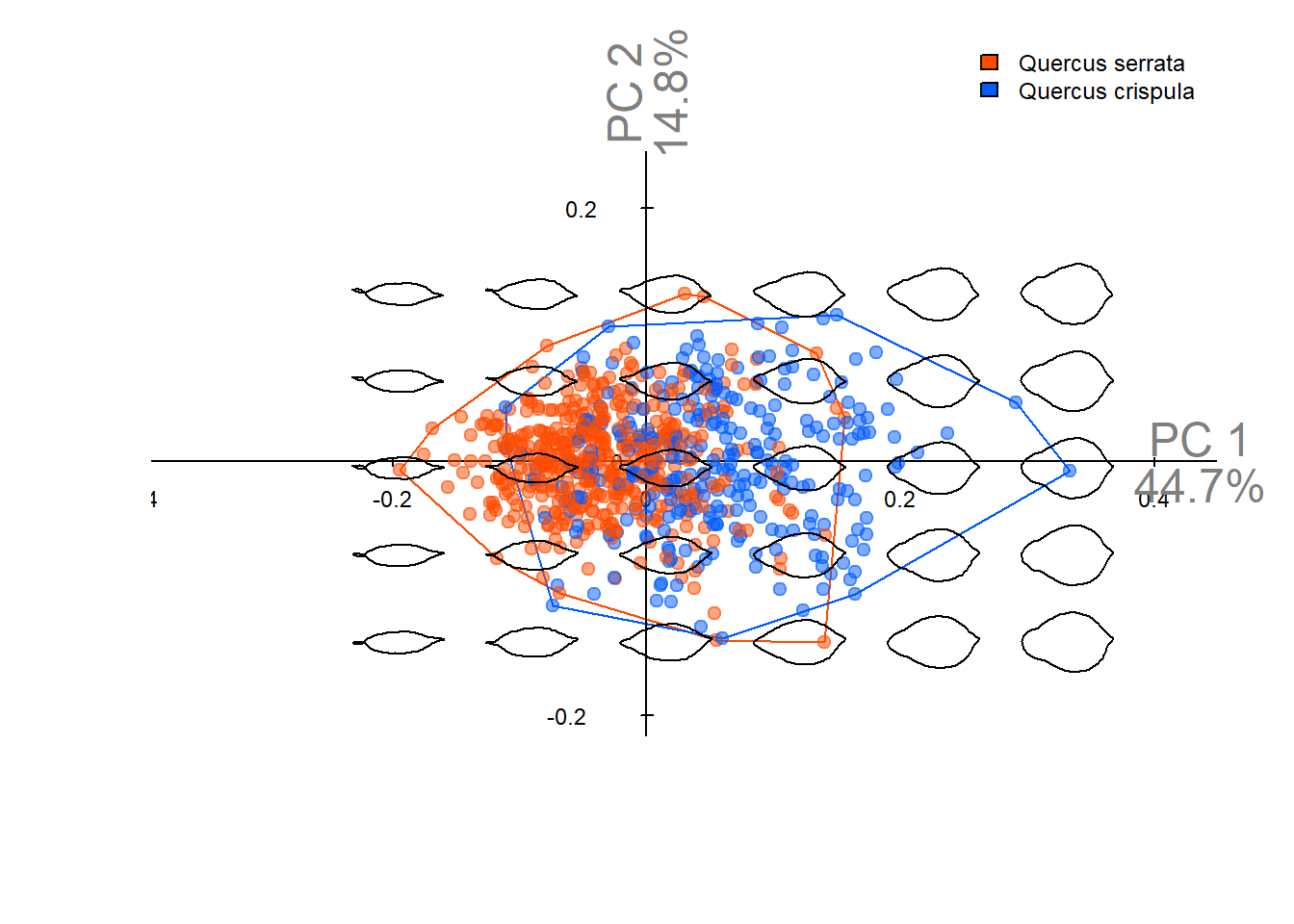
Output figures in this notebook are used in **Figure 4A-C** of the manuscript.
## Setup & preprocessing
```{r}
#| label: setup
library(tools)
library(Momocs)
source("R/geometry.R")
source("R/normalization.R")
source("R/pca_plot.R")
# color palette for colorblind-friendly visualization
palette("Okabe-Ito")
# Quercus serrata and Q. crispula colors
my_palette <- pal_manual(c("#FF4B00", "#005AFF"), transp = 0)
# set the data directory from environment variable (.Renviron)
data_dir <- Sys.getenv("PROJECT_DATA_DIR")
```
Load the species names from the CSV file.
This file contains three columns: `id`, `konara`, and `mizunara`. The `id` column corresponds to the sample ID, while the `konara` and `mizunara` columns contain binary values indicating the species classification. We will determine the species name based on which column has a value of 1 for each sample.
::: callout-note
## Species name
`konara` means Quercus serrata, and `mizunara` means Quercus crispula. Both are standard Japanese common names used in the original dataset metadata.
:::
```{r}
#| label: species_name
df_sp_name <- read.csv(file.path(data_dir, "data/species_name.csv"))
df_sp_name$species <- ifelse(
df_sp_name$konara < df_sp_name$mizunara,
"Quercus crispula",
"Quercus serrata"
)
df_sp_name$species <- factor(
df_sp_name$species,
levels = c("Quercus serrata", "Quercus crispula")
)
```
Load contours from the CSV files in the `data/contour_quercus/` directory.
```{r}
#| label: load_contours
file_paths_xy <- list.files(
file.path(data_dir, "data/contour_quercus/"),
full.names = TRUE,
pattern = "\\.csv$"
)
xy_list <- lapply(file_paths_xy, function(fp) {
#cat("File:", fp, "\n")
df <- read.csv(fp)
df <- df[, c("x", "y")]
df <- as.matrix(df)
storage.mode(df) <- "double"
# If the contour is not closed, append the first point to the end.
if (!all(df[1, ] == df[nrow(df), ])) {
df <- rbind(df, df[1, ])
}
return(df)
})
names(xy_list) <- file_path_sans_ext(basename(file_paths_xy))
```
Rotate contour to randomize the initial direction.
```{r}
#| label: rotate_contours
set.seed(123) # set.seed for reproducibility
angles <- sample(0:360, length(xy_list), replace = TRUE) # rotation angles in degrees
# rotate contours
xy_list_rotated <- mapply(
function(mat, ang) rotate_xy_centered(mat, ang),
xy_list,
angles,
SIMPLIFY = FALSE
)
```
To evaluate the normalization methods without bias from the contour’s initial point, we randomize the starting point of each contour.
This is necessary because the contours obtained with [LeafContourEFD](https://github.com/maple60/leaf-contour-efd) already have a standardized starting point.
To randomize the starting point, we reorder the points in each contour by selecting a starting point at random and rearranging the sequence accordingly.
```{r}
#| label: randomize_starting_point
set.seed(123)
xy_list_rotated_reordered <- lapply(xy_list_rotated, function(xy_rotated) {
# rotation angles in degrees
idx_start <- sample(seq_len(nrow(xy_rotated)), 1)
xy_rotated_reordered <- xy_rotated[
c(idx_start:nrow(xy_rotated), 1:(idx_start - 1)),
]
return(xy_rotated_reordered)
})
```
Species names corresponding to individual IDs.
```{r}
#| label: prepare_ids_species
id_name <- file_path_sans_ext(basename(file_paths_xy)) # IDs with suffixes
# individual IDs without suffixes
id_name_head <- lapply(id_name, function(x) {
parts <- unlist(strsplit(x, "_"))
return(parts[1])
})
id_name_head <- unlist(id_name_head)
# species names corresponding to individual IDs
sp_name <- df_sp_name$species[match(id_name_head, df_sp_name$id)]
```
## PCA using conventional normalization
Principal component analysis (PCA) using normalized EFD coefficients (conventional method [@kuhl1982]).\
Normalization is performed using [Momocs](https://momx.github.io/Momocs/) [@bonhomme2014].
```{r}
#| label: pca_efd_normalized
nb_h <- 35 # Number of harmonics
out <- Out(
xy_list_rotated_reordered,
fac = data.frame(species = factor(sp_name))
) # convert to Momocs Out object
# EFD with normalization
ef_norm <- efourier(out, nb.h = nb_h, norm = TRUE, start = FALSE)
pca_norm <- PCA(ef_norm, center = TRUE, scale. = FALSE) # PCA with Momocs
idx_max <- which.max(pca_norm$x[, 2])
idx_min <- which.min(pca_norm$x[, 2])
pca_plot(pca_norm) # visualize PCA results
```
```{r}
#| label: pca_efd_normalized_save
#| eval: false
#| code-fold: true
#| code-summary: "Save the plot as SVG file"
save_dir <- "results"
dir.create(save_dir, showWarnings = FALSE, recursive = TRUE)
save_file_name <- "fig4a_pca_efd_normalized_quercus.svg"
save_file_path <- file.path(save_dir, save_file_name)
svg(
save_file_path,
bg = "transparent"
)
pca_plot(pca_norm)
dev.off()
```
## PCA using true EFD normalization
Principal component analysis (PCA) using coefficients obtained by true EFD normalization.\
Normalization is performed using the `efourier_true_norm()` function implemented in this project (see [`R/normalization.R`](https://github.com/maple60/leaf-contour-efd-analysis/blob/f04a8cd81357d5195f1da90fc051b169971a3abd/R/normalization.R) for details).
```{r}
#| label: pca_true_efd_normalized
nb_h <- 35 # Number of harmonics
# convert to Momocs Out object
out <- Out(
xy_list_rotated_reordered,
fac = data.frame(species = factor(sp_name))
)
# oriented true EFD normalization
ef_true_norm <- efourier_true_norm(
out,
nb_h = nb_h,
x_sy = TRUE,
y_sy = TRUE,
skip_rotation = FALSE
)
# PCA with Momocs
pca_true_norm <- PCA(ef_true_norm, center = TRUE, scale. = FALSE)
# flip PC axes for better visualization
pca_true_norm <- flip_PCaxes(pca_true_norm, 1)
pca_plot(pca_true_norm)
```
```{r}
#| label: pca_true_efd_normalized_save
#| eval: false
#| code-fold: true
#| code-summary: "Save the plot as SVG file"
save_dir <- "results"
dir.create(save_dir, showWarnings = FALSE, recursive = TRUE)
save_file_name <- "fig4b_pca_true_efd_normalized.svg"
save_file_path <- file.path(save_dir, save_file_name)
svg(
save_file_path,
bg = "transparent"
)
pca_plot(pca_true_norm)
dev.off()
```
## PCA using oriented true EFD normalization
Contours obtained from [LeafContourEFD](https://github.com/maple60/leaf-contour-efd) already have standardized starting points and rotation.
Therefore, we apply only the x-axis symmetry correction step from **true EFD normalization** to implement oriented **true EFD normalization**.
Although the same results can be obtained from the Fourier coefficients exported by [LeafContourEFD](https://github.com/maple60/leaf-contour-efd), we use [Momocs](https://momx.github.io/Momocs/) here so we can easily visualize reconstructed contour changes associated with principal component score shifts.
In a future update, we plan to provide a workflow that does not depend on [Momocs](https://momx.github.io/Momocs/).
```{r}
#| label: pca_oriented_true_efd_normalized
nb_h <- 35 # Number of harmonics
out <- Out(xy_list, fac = data.frame(species = factor(sp_name))) # convert to Momocs Out object
# oriented true EFD normalization
ef_true_norm <- efourier_true_norm(
out,
nb_h = nb_h,
x_sy = TRUE,
y_sy = FALSE,
skip_rotation = TRUE
)
# PCA with Momocs
pca_true_norm <- PCA(ef_true_norm, center = TRUE, scale. = FALSE)
```
Visualize PCA results.
```{r}
#| label: pca_oriented_true_efd_normalized_plot
pca_plot(pca_true_norm)
```
```{r}
#| label: pca_oriented_true_efd_normalized_save
#| eval: false
#| code-fold: true
#| code-summary: "Save the plot as SVG file"
save_dir <- "results"
dir.create(save_dir, showWarnings = FALSE, recursive = TRUE)
save_file_name <- "fig4c_pca_oriented_true_efd_normalized.svg"
save_file_path <- file.path(save_dir, save_file_name)
svg(
save_file_path,
bg = "transparent"
)
pca_plot(pca_true_norm)
dev.off()
```
## References
::: {#refs}
:::